用 Thomson Sampling 对旋转矩阵采样
本文说明用 Thomson 球面采样思想对三维旋转矩阵R∈SO3的采样,并解释为什么它比“直接对三个欧拉角均匀采样”更适合生成近似均匀的旋转样本。代码在本文末尾。
本文说明用 Thomson 球面采样思想对三维旋转矩阵 R∈SO(3)R \in SO(3)R∈SO(3) 的采样,并解释为什么它比“直接对三个欧拉角均匀采样”更适合生成近似均匀的旋转样本。 代码ParticleSampleSphere.m在本文末尾。
1. 问题背景
三维旋转矩阵构成特殊正交群:
SO(3)={R∈R3×3∣RTR=I, det(R)=1}. SO(3) = \{ R \in \mathbb{R}^{3 \times 3} \mid R^{T} R = I,\ \det(R) = 1 \}. SO(3)={R∈R3×3∣RTR=I, det(R)=1}.
虽然一个旋转矩阵可以用三个参数表示,例如欧拉角、滚转-俯仰-偏航角,但 SO(3)SO(3)SO(3) 不是普通的三维欧氏空间。直接令三个角度独立均匀分布:
ϕ, θ, ψ∼U(⋅) \phi,\ \theta,\ \psi \sim U(\cdot) ϕ, θ, ψ∼U(⋅)
通常不会得到 SO(3)SO(3)SO(3) 上均匀分布的旋转。原因是欧拉角参数化存在奇异性和非均匀体积元素。例如 ZYX 欧拉角的 Haar 测度中间角带有雅可比权重,形式上类似:
dμ(R)∝cos(θ) dϕ dθ dψ. d\mu(R) \propto \cos(\theta)\, d\phi\, d\theta\, d\psi. dμ(R)∝cos(θ)dϕdθdψ.
因此,若直接对 θ\thetaθ 均匀采样,就会破坏旋转空间上的均匀性。一些采样点会相互靠得很近,另一些区域会比较稀疏,导致搜索或匹配时出现冗余样本。
2. 用四元数表示旋转
单位四元数:
q=(w,x,y,z)T,∥q∥=1 q = (w, x, y, z)^{T}, \quad \lVert q \rVert = 1 q=(w,x,y,z)T,∥q∥=1
位于四维空间中的三维单位球面:
S3={q∈R4∣∥q∥=1}. S^{3} = \{ q \in \mathbb{R}^{4} \mid \lVert q \rVert = 1 \}. S3={q∈R4∣∥q∥=1}.
每个单位四元数可以表示一个三维旋转。其对应旋转矩阵为:
R(q)=[1−2y2−2z22xy−2zw2xz+2yw2xy+2zw1−2x2−2z22yz−2xw2xz−2yw2yz+2xw1−2x2−2y2]. R(q) = \begin{bmatrix} 1 - 2y^{2} - 2z^{2} & 2xy - 2zw & 2xz + 2yw \\ 2xy + 2zw & 1 - 2x^{2} - 2z^{2} & 2yz - 2xw \\ 2xz - 2yw & 2yz + 2xw & 1 - 2x^{2} - 2y^{2} \end{bmatrix}. R(q)= 1−2y2−2z22xy+2zw2xz−2yw2xy−2zw1−2x2−2z22yz+2xw2xz+2yw2yz−2xw1−2x2−2y2 .
需要注意,四元数是旋转群 SO(3)SO(3)SO(3) 的双覆盖:
q和−q q \quad \text{和} \quad -q q和−q
表示同一个旋转。因此在比较两个旋转对应的四元数时,需要使用绝对内积。
3. 旋转之间的距离
两个旋转矩阵 Ri, RjR_i,\ R_jRi, Rj 的相对旋转角为:
dSO(3)(Ri,Rj)=arccos (tr(RiTRj)−12). d_{SO(3)}(R_i, R_j)= \arccos \! \left( \frac{\mathrm{tr}(R_i^{T} R_j) - 1}{2} \right). dSO(3)(Ri,Rj)=arccos(2tr(RiTRj)−1).
若它们对应单位四元数 qi, qjq_i,\ q_jqi, qj ,则同一个距离可以写为:
dij=2arccos(∣qiTqj∣),dij∈[0,π]. d_{ij}= 2 \arccos( \lvert q_i^{T} q_j \rvert ), \quad d_{ij} \in [0, \pi]. dij=2arccos(∣qiTqj∣),dij∈[0,π].
这里的 ∣qiTqj∣\lvert q_i^{T} q_j \rvert∣qiTqj∣ 正是为了处理 qqq 与 −q-q−q 表示同一旋转的问题。
4. Thomson Sampling 的能量模型
本项目的 ParticleSampleSphere.m 通过最小化带电粒子系统的 Riesz sss-energy,在球面上生成近似均匀的点。对旋转采样时,可以把每个单位四元数 qiq_iqi 看作 S3S^{3}S3 上的一个同号带电粒子。
给定 NNN 个旋转样本:
Q={q1,q2,…,qN},qi∈S3, Q = \{ q_1, q_2, \dots, q_N \}, \quad q_i \in S^{3}, Q={q1,q2,…,qN},qi∈S3,
定义总能量:
E(Q)=∑1≤i<j≤N1(dij+ϵ)s. E(Q)= \sum_{1 \le i < j \le N} \frac{1}{(d_{ij} + \epsilon)^{s}}. E(Q)=1≤i<j≤N∑(dij+ϵ)s1.
其中:
- s>0s > 0s>0 是 Riesz 能量参数。
- s=1s = 1s=1 对应经典 Thomson 问题。
- ϵ\epsilonϵ 是很小的数,用于避免数值除零。
- dij=2arccos(∣qiTqj∣)d_{ij} = 2 \arccos( \lvert q_i^{T} q_j \rvert )dij=2arccos(∣qiTqj∣) 是两个旋转之间的测地距离。
当两个旋转很接近时, dijd_{ij}dij 很小,能量项很大,于是优化会把它们推开。最小化总能量后,点集会趋向于在旋转空间中均匀展开。
5. 梯度下降更新
记:
cij=qiTqj,aij=∣cij∣,dij=2arccos(aij). c_{ij} = q_i^{T} q_j, \quad a_{ij} = \lvert c_{ij} \rvert, \quad d_{ij} = 2 \arccos(a_{ij}). cij=qiTqj,aij=∣cij∣,dij=2arccos(aij).
单个能量项为:
Eij=1(dij+ϵ)s. E_{ij} = \frac{1}{(d_{ij} + \epsilon)^{s}}. Eij=(dij+ϵ)s1.
对 qiq_iqi 的欧氏梯度可写为:
∇qiE=∑j≠i2s sign(cij) qj(dij+ϵ)s+11−cij2+ϵ. \nabla_{q_i} E= \sum_{j \ne i} \frac{2 s \, \mathrm{sign}(c_{ij}) \, q_j} {(d_{ij} + \epsilon)^{s+1} \sqrt{1 - c_{ij}^{2} + \epsilon}}. ∇qiE=j=i∑(dij+ϵ)s+11−cij2+ϵ2ssign(cij)qj.
但 qiq_iqi 必须始终留在单位球面 S3S^{3}S3 上,所以不能直接用欧氏梯度更新。需要先把梯度投影到 S3S^{3}S3 在 qiq_iqi 处的切空间:
gi⊥=gi−(giTqi)qi. g_i^{\perp}= g_i - (g_i^{T} q_i) q_i. gi⊥=gi−(giTqi)qi.
然后沿负梯度方向更新并重新归一化:
qinew=qi−αgi⊥∥qi−αgi⊥∥. q_i^{\text{new}}= \frac{q_i - \alpha g_i^{\perp}} {\lVert q_i - \alpha g_i^{\perp} \rVert}. qinew=∥qi−αgi⊥∥qi−αgi⊥.
脚本中的实现还加入了简单的回溯线搜索:如果某一步更新后总能量没有下降,就缩小步长再试。
6. 完整流程
用 Thomson sampling 生成旋转矩阵样本的流程如下:
- 随机生成 NNN 个四维向量。
- 将每个向量归一化到 S3S^{3}S3 ,得到初始单位四元数。
- 计算所有样本两两之间的旋转距离 dijd_{ij}dij 。
- 计算 Riesz sss-energy。
- 计算能量梯度。
- 将梯度投影到 S3S^{3}S3 的切空间。
- 更新四元数并重新归一化。
- 重复迭代,直到能量变化很小或达到最大迭代次数。
- 将最终四元数转换成旋转矩阵。
7. 与欧拉角均匀采样的对比指标
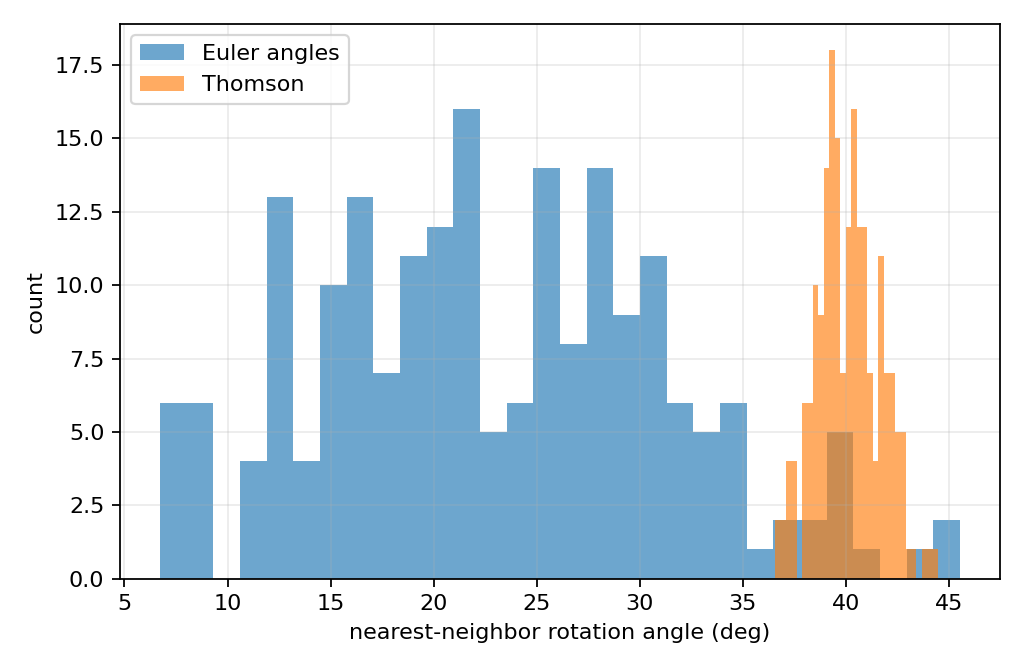
脚本 thomson_rotation_sampling.py 使用最近邻旋转距离作为主要对比指标。
对每个样本 RiR_iRi ,计算它与其他样本之间的最小旋转角:
δi=minj≠idSO(3)(Ri,Rj). \delta_i= \min_{j \ne i} d_{SO(3)}(R_i, R_j). δi=j=imindSO(3)(Ri,Rj).
如果采样更均匀,则最近邻距离通常具有这些特征:
- 最小值更大,说明没有非常近的冗余样本。
- 标准差更小,说明样本间距更一致。
- 变异系数 std(δi)/mean(δi)\mathrm{std}(\delta_i) / \mathrm{mean}(\delta_i)std(δi)/mean(δi) 更小。
- Riesz 能量更低,说明近距离冲突更少。
欧拉角均匀采样一般会出现更小的最近邻距离和更大的距离波动。
8. 运行脚本
基本运行:
python3 thomson_rotation_sampling.py --n 200 --iterations 800 --seed 7
输出文件默认以 rotation_sampling 为前缀,保存一张 PNG:
rotation_sampling_comparison.png
9. thomson_rotation_sampling.py 源代码
from __future__ import annotations
import argparse
import struct
import zlib
from dataclasses import dataclass
from pathlib import Path
import numpy as np
EPS = 1e-12
@dataclass
class ThomsonResult:
quaternions: np.ndarray
energies: list[float]
iterations: int
def normalize_rows(x: np.ndarray) -> np.ndarray:
norms = np.linalg.norm(x, axis=1, keepdims=True)
return x / np.maximum(norms, EPS)
def canonicalize_quaternions(q: np.ndarray) -> np.ndarray:
"""Choose a stable representative for q ~ -q."""
q = q.copy()
mask = q[:, 0] < 0
q[mask] *= -1.0
return q
def random_quaternions(n: int, rng: np.random.Generator) -> np.ndarray:
q = rng.normal(size=(n, 4))
return normalize_rows(q)
def quaternion_multiply(a: np.ndarray, b: np.ndarray) -> np.ndarray:
aw, ax, ay, az = np.moveaxis(a, -1, 0)
bw, bx, by, bz = np.moveaxis(b, -1, 0)
return np.stack(
[
aw * bw - ax * bx - ay * by - az * bz,
aw * bx + ax * bw + ay * bz - az * by,
aw * by - ax * bz + ay * bw + az * bx,
aw * bz + ax * by - ay * bx + az * bw,
],
axis=-1,
)
def euler_zyx_quaternions(n: int, rng: np.random.Generator) -> np.ndarray:
"""Naive independent uniform yaw-pitch-roll sampling.
Uses R = Rz(yaw) Ry(pitch) Rx(roll). Pitch is sampled in [-pi/2, pi/2]
and yaw/roll in [-pi, pi]. This is a common "uniform three angles"
baseline, but it is not uniform on SO(3).
"""
yaw = rng.uniform(-np.pi, np.pi, size=n)
pitch = rng.uniform(-np.pi / 2.0, np.pi / 2.0, size=n)
roll = rng.uniform(-np.pi, np.pi, size=n)
zeros = np.zeros(n)
cy = np.cos(yaw / 2.0)
sy = np.sin(yaw / 2.0)
qz = np.stack([cy, zeros, zeros, sy], axis=1)
cp = np.cos(pitch / 2.0)
sp = np.sin(pitch / 2.0)
qy = np.stack([cp, zeros, sp, zeros], axis=1)
cr = np.cos(roll / 2.0)
sr = np.sin(roll / 2.0)
qx = np.stack([cr, sr, zeros, zeros], axis=1)
q = quaternion_multiply(quaternion_multiply(qz, qy), qx)
return canonicalize_quaternions(normalize_rows(q))
def quaternions_to_rotation_matrices(q: np.ndarray) -> np.ndarray:
q = normalize_rows(q)
w, x, y, z = q.T
rotations = np.empty((q.shape[0], 3, 3), dtype=float)
rotations[:, 0, 0] = 1.0 - 2.0 * (y * y + z * z)
rotations[:, 0, 1] = 2.0 * (x * y - z * w)
rotations[:, 0, 2] = 2.0 * (x * z + y * w)
rotations[:, 1, 0] = 2.0 * (x * y + z * w)
rotations[:, 1, 1] = 1.0 - 2.0 * (x * x + z * z)
rotations[:, 1, 2] = 2.0 * (y * z - x * w)
rotations[:, 2, 0] = 2.0 * (x * z - y * w)
rotations[:, 2, 1] = 2.0 * (y * z + x * w)
rotations[:, 2, 2] = 1.0 - 2.0 * (x * x + y * y)
return rotations
def pairwise_rotation_angles(q: np.ndarray) -> np.ndarray:
q = normalize_rows(q)
dots = np.abs(q @ q.T)
dots = np.clip(dots, 0.0, 1.0)
return 2.0 * np.arccos(dots)
def nearest_neighbor_angles(q: np.ndarray) -> np.ndarray:
angles = pairwise_rotation_angles(q)
np.fill_diagonal(angles, np.inf)
return np.min(angles, axis=1)
def thomson_energy(q: np.ndarray, s: float = 1.0, eps: float = EPS) -> float:
angles = pairwise_rotation_angles(q)
i_upper = np.triu_indices(q.shape[0], k=1)
d = np.maximum(angles[i_upper], eps)
return float(np.sum(1.0 / (d**s)))
def thomson_gradient(q: np.ndarray, s: float = 1.0, eps: float = EPS) -> np.ndarray:
"""Projected gradient of sum 1 / theta_ij^s on S^3 / {q ~ -q}."""
dots = q @ q.T
signs = np.sign(dots)
abs_dots = np.clip(np.abs(dots), 0.0, 1.0 - 1e-10)
angles = 2.0 * np.arccos(abs_dots)
denom = (np.maximum(angles, eps) ** (s + 1.0)) * np.sqrt(
np.maximum(1.0 - abs_dots * abs_dots, eps)
)
weights = 2.0 * s * signs / denom
np.fill_diagonal(weights, 0.0)
grad = weights @ q
radial = np.sum(grad * q, axis=1, keepdims=True) * q
return grad - radial
def thomson_sample_rotations(
n: int,
iterations: int,
seed: int,
s: float,
step_size: float,
etol: float,
dtol: float,
report_every: int,
quiet: bool,
) -> ThomsonResult:
rng = np.random.default_rng(seed)
q = random_quaternions(n, rng)
energy = thomson_energy(q, s=s)
energies = [energy]
step = step_size
min_step = 1e-8
if not quiet:
print(f"Initial Thomson energy: {energy:.6f}")
completed = 0
for iteration in range(1, iterations + 1):
grad = thomson_gradient(q, s=s)
grad_norm = np.linalg.norm(grad, axis=1, keepdims=True)
direction = grad / np.maximum(grad_norm, EPS)
accepted = False
trial_step = step
displacement = 0.0
candidate_energy = energy
candidate = q
for _ in range(30):
candidate = normalize_rows(q - trial_step * direction)
candidate_energy = thomson_energy(candidate, s=s)
if candidate_energy < energy:
dots = np.abs(np.sum(q * candidate, axis=1))
dots = np.clip(dots, 0.0, 1.0)
displacement = float(np.max(2.0 * np.arccos(dots)))
accepted = True
break
trial_step *= 0.5
if not accepted:
step = trial_step
if step < min_step:
if not quiet:
print("Stopped: line search step became too small.")
break
continue
q = candidate
energy_drop = energy - candidate_energy
energy = candidate_energy
energies.append(energy)
step = min(trial_step * 1.03, 0.35)
completed = iteration
if not quiet and (iteration == 1 or iteration % report_every == 0):
print(
f"iter={iteration:5d} energy={energy:.6f} "
f"dE={energy_drop:.3e} max_step_deg={np.degrees(displacement):.5f}"
)
if iteration > 10 and energy_drop < etol and displacement < dtol:
if not quiet:
print("Stopped: energy and displacement tolerances reached.")
break
return ThomsonResult(
quaternions=canonicalize_quaternions(q),
energies=energies,
iterations=completed,
)
def summarize(name: str, q: np.ndarray, s: float) -> dict[str, float | str | int]:
nn = np.degrees(nearest_neighbor_angles(q))
mean = float(np.mean(nn))
std = float(np.std(nn))
return {
"method": name,
"n": int(q.shape[0]),
"energy": thomson_energy(q, s=s),
"nn_min_deg": float(np.min(nn)),
"nn_p05_deg": float(np.percentile(nn, 5)),
"nn_mean_deg": mean,
"nn_median_deg": float(np.median(nn)),
"nn_p95_deg": float(np.percentile(nn, 95)),
"nn_max_deg": float(np.max(nn)),
"nn_std_deg": std,
"nn_cv": float(std / mean) if mean > 0.0 else float("nan"),
}
def print_summary(rows: list[dict[str, float | str | int]]) -> None:
print("\nComparison summary")
print(
"method energy min(deg) mean(deg) std(deg) cv p05(deg)"
)
for row in rows:
print(
f"{row['method']:<10} "
f"{row['energy']:>12.4f} "
f"{row['nn_min_deg']:>9.3f} "
f"{row['nn_mean_deg']:>10.3f} "
f"{row['nn_std_deg']:>9.3f} "
f"{row['nn_cv']:>8.3f} "
f"{row['nn_p05_deg']:>9.3f}"
)
def write_png(path: Path, image: np.ndarray) -> None:
"""Write an RGB uint8 image as a PNG using only the standard library."""
path.parent.mkdir(parents=True, exist_ok=True)
if image.dtype != np.uint8 or image.ndim != 3 or image.shape[2] != 3:
raise ValueError("image must be an RGB uint8 array")
height, width, _ = image.shape
raw = bytearray()
for row in image:
raw.append(0)
raw.extend(row.tobytes())
def chunk(tag: bytes, data: bytes) -> bytes:
crc = zlib.crc32(tag + data) & 0xFFFFFFFF
return struct.pack(">I", len(data)) + tag + data + struct.pack(">I", crc)
png = [
b"\x89PNG\r\n\x1a\n",
chunk(b"IHDR", struct.pack(">IIBBBBB", width, height, 8, 2, 0, 0, 0)),
chunk(b"IDAT", zlib.compress(bytes(raw), level=9)),
chunk(b"IEND", b""),
]
path.write_bytes(b"".join(png))
def blend_rect(
image: np.ndarray,
x0: int,
y0: int,
x1: int,
y1: int,
color: tuple[int, int, int],
alpha: float = 1.0,
) -> None:
height, width, _ = image.shape
xa = max(0, min(width, int(round(x0))))
xb = max(0, min(width, int(round(x1))))
ya = max(0, min(height, int(round(y0))))
yb = max(0, min(height, int(round(y1))))
if xa >= xb or ya >= yb:
return
base = image[ya:yb, xa:xb].astype(float)
overlay = np.array(color, dtype=float)
image[ya:yb, xa:xb] = np.round(base * (1.0 - alpha) + overlay * alpha).astype(
np.uint8
)
def draw_fallback_histogram_png(
path: Path,
thomson_nn: np.ndarray,
euler_nn: np.ndarray,
bins: int = 30,
) -> None:
"""Small dependency-free histogram renderer used when matplotlib is absent."""
width, height = 1000, 620
left, right, top, bottom = 90, 35, 45, 80
plot_w = width - left - right
plot_h = height - top - bottom
image = np.full((height, width, 3), 255, dtype=np.uint8)
max_angle = max(float(np.max(thomson_nn)), float(np.max(euler_nn)), 1.0)
edges = np.linspace(0.0, max_angle * 1.05, bins + 1)
th_counts, _ = np.histogram(thomson_nn, bins=edges)
eu_counts, _ = np.histogram(euler_nn, bins=edges)
max_count = max(int(np.max(th_counts)), int(np.max(eu_counts)), 1)
axis = (70, 70, 70)
grid = (225, 225, 225)
thomson_color = (39, 119, 180)
euler_color = (221, 126, 55)
for i in range(6):
y = top + plot_h - i * plot_h / 5
blend_rect(image, left, y, width - right, y + 1, grid, 1.0)
blend_rect(image, left, top, left + 2, height - bottom, axis, 1.0)
blend_rect(image, left, height - bottom, width - right, height - bottom + 2, axis, 1.0)
bin_w = plot_w / bins
for i in range(bins):
x0 = left + i * bin_w
x1 = left + (i + 1) * bin_w
th_h = plot_h * th_counts[i] / max_count
eu_h = plot_h * eu_counts[i] / max_count
blend_rect(
image,
x0 + 2,
height - bottom - eu_h,
x1 - 1,
height - bottom,
euler_color,
0.62,
)
blend_rect(
image,
x0 + 2,
height - bottom - th_h,
x1 - 1,
height - bottom,
thomson_color,
0.62,
)
# Lightweight legend swatches; matplotlib is used when full labels are available.
blend_rect(image, width - right - 210, top + 18, width - right - 175, top + 38, euler_color, 0.8)
blend_rect(image, width - right - 210, top + 48, width - right - 175, top + 68, thomson_color, 0.8)
write_png(path, image)
def plot_nearest_neighbor_histogram(prefix: Path, thomson_q: np.ndarray, euler_q: np.ndarray) -> Path:
out = prefix.with_name(prefix.name + "_comparison.png")
thomson_nn = np.degrees(nearest_neighbor_angles(thomson_q))
euler_nn = np.degrees(nearest_neighbor_angles(euler_q))
try:
import matplotlib.pyplot as plt
except ModuleNotFoundError:
draw_fallback_histogram_png(out, thomson_nn, euler_nn)
print("matplotlib is not installed; wrote a simple PNG histogram instead.")
return out
prefix.parent.mkdir(parents=True, exist_ok=True)
fig, ax = plt.subplots(figsize=(6.4, 4.2))
ax.hist(euler_nn, bins=30, alpha=0.65, label="Euler angles")
ax.hist(thomson_nn, bins=30, alpha=0.65, label="Thomson")
ax.set_xlabel("nearest-neighbor rotation angle (deg)")
ax.set_ylabel("count")
ax.legend()
ax.grid(alpha=0.25)
fig.tight_layout()
fig.savefig(out, dpi=160)
plt.close(fig)
return out
def parse_args() -> argparse.Namespace:
parser = argparse.ArgumentParser(
description="Sample SO(3) with Thomson sampling and compare against naive uniform Euler angles."
)
parser.add_argument("--n", type=int, default=200, help="number of rotations")
parser.add_argument("--iterations", type=int, default=800, help="maximum optimization iterations")
parser.add_argument("--seed", type=int, default=7, help="random seed")
parser.add_argument("--s-energy", type=float, default=1.0, help="Riesz s-energy exponent")
parser.add_argument("--step-size", type=float, default=0.05, help="initial tangent step size")
parser.add_argument("--etol", type=float, default=1e-10, help="energy-drop tolerance")
parser.add_argument("--dtol", type=float, default=1e-8, help="max displacement tolerance in radians")
parser.add_argument("--report-every", type=int, default=100, help="progress print interval")
parser.add_argument(
"--output-prefix",
type=Path,
default=Path("rotation_sampling"),
help="prefix for the output PNG file",
)
parser.add_argument("--quiet", action="store_true", help="suppress optimization progress")
return parser.parse_args()
def main() -> None:
args = parse_args()
if args.n < 2:
raise ValueError("--n must be at least 2")
if args.iterations < 1:
raise ValueError("--iterations must be at least 1")
if args.s_energy <= 0.0:
raise ValueError("--s-energy must be positive")
result = thomson_sample_rotations(
n=args.n,
iterations=args.iterations,
seed=args.seed,
s=args.s_energy,
step_size=args.step_size,
etol=args.etol,
dtol=args.dtol,
report_every=max(args.report_every, 1),
quiet=args.quiet,
)
rng = np.random.default_rng(args.seed + 1)
euler_q = euler_zyx_quaternions(args.n, rng)
rows = [
summarize("Thomson", result.quaternions, s=args.s_energy),
summarize("Euler", euler_q, s=args.s_energy),
]
print_summary(rows)
plot_path = plot_nearest_neighbor_histogram(
args.output_prefix, result.quaternions, euler_q
)
print("\nSaved output")
print(f"- {plot_path}")
print(f"\nAccepted Thomson iterations: {result.iterations}")
if __name__ == "__main__":
main()

openEuler 是由开放原子开源基金会孵化的全场景开源操作系统项目,面向数字基础设施四大核心场景(服务器、云计算、边缘计算、嵌入式),全面支持 ARM、x86、RISC-V、loongArch、PowerPC、SW-64 等多样性计算架构
更多推荐
 已为社区贡献1条内容
已为社区贡献1条内容







所有评论(0)